Cool tools: understanding communication between cells
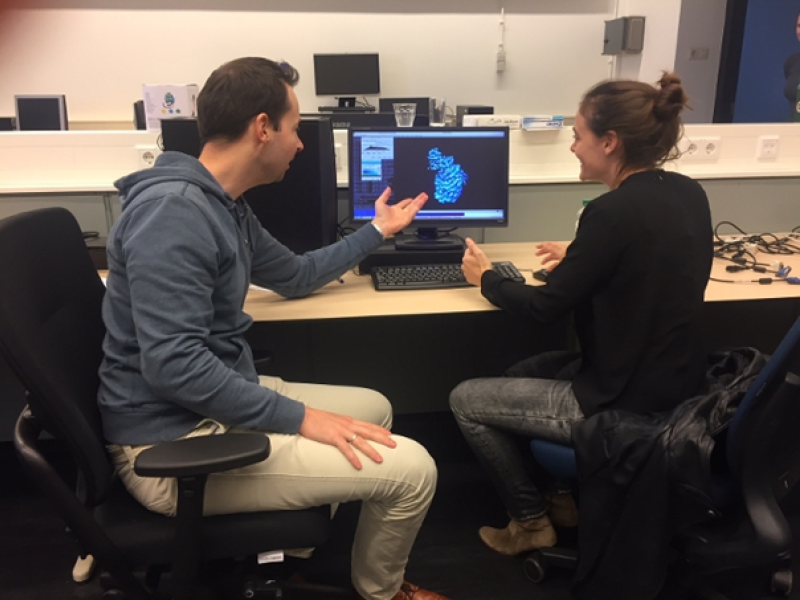
Chemists Bert Janssen and Dimphna Meijer of Utrecht University research the proteins that make communication between brain cells possible. These minuscule particles are made visible with a cryo-electron microscope. A million images are then combined into a three-dimensional image with the help of the Dutch National Supercomputer.
Conversation between brain cells
The communication between brain cells is essential for our brain's functioning and development. This 'conversation' is usually facilitated by proteins. Dr Bert Janssen's research group Crystal and Structural Chemistry at Utrecht University examines how these protein systems initiate and convert signals between our brain cells.
How the healthy brain works
"We want to gain a fundamental understanding of how the brain works at the molecular level," Dimphna Meijer, a post-doctoral researcher at Janssen's group, says. "Rather than study what goes wrong in case of a disease, we examine why things go well in a healthy brain. This will ultimately allow us to better understand what happens when things go wrong."
1000 times smaller than a hair
The point of interaction between two neurons (nerve cells) is called a synapse. Bert Janssen and Dimphna Meijer have mapped a specific protein that ensures that these synapses stick together: Teneurin. "This protein is 20 nanometres long," Janssen says. "That's 1000 times smaller than the diameter of a hair. You can't even see it with a normal microscope. We use an electron microscope."
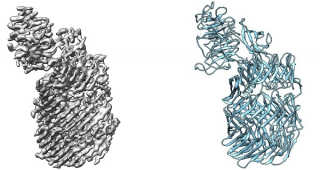
The teneurin protein
Frozen proteins
Cryo-electron microscopy is a relatively new and revolutionary technology (according to Nature) that was awarded a Nobel Prize in 2017. This technique fires electron beams at frozen protein samples to make the structure of the biomolecules visible. The computer then combines the resulting images taken from different angles into 3D imagery. This also involves the Relion software developed by Dutchman Sjors Scheres, who completed his PhD at Utrecht University and is now leading a research group in Cambridge.
2.1 million core hours
Converting the images takes a lot of computer power. The chemists Janssen en Meijer work with a GPU cluster on the Cartesius National Supercomputer. The supercomputer uses many processor cores simultaneously. On a normal laptop with 4 cores, these calculations would take almost 60 years. The research group has been allocated significant capacity: 2.1 million core hours, of which a third has already been used in one year. Graphics processing units (GPUs) can perform the necessary computing tasks much faster than traditional core processing units (CPUs). Meijer: "SURFsara has helped us to make optimal use of our algorithms on the GPU cluster."
11 terabytes of data
"To get a three-dimensional image of the protein at a high resolution, we need a million images which are subsequently combined with each other. Our dataset started with 11 terabytes of data. This all had to be transferred to Cartesius, which took 3 days. During the processing, which takes only a few hours, the amount of data doubles again. The final 3D image is then reduced to something that can be put on a USB stick: 200 MB.”
New questions
The 3D image of the Teneurin protein has resulted in a first publication for this project. Now that the researchers have a good picture of the protein, they can study it further. Janssen: "The most important conclusion is that Teneurin looks very different from all the traditional cell adhesion molecules, which are the proteins involved in connecting cells together. The odd thing is that its structure is extremely similar to a bacterial protein. Now that we have a detailed picture of how it is composed, what the functional units look like and where they are in relation to each other, we can move on. We want to understand how the protein works. We also do not know yet why it looks so different. And as usual, this discovery raises many other new questions."
Publication
Structures of Teneurin adhesion receptors reveal an ancient fold for cell-cell interaction
More information
In this video Bert Jansen talks about his research

