“It is quite extraordinary that such a project is taking place here in the Netherlands”
Digging into a genetic goldmine
MinE is an international project that aims to identify the genetic causes of amyotrophic lateral sclerosis (ALS). ALS is a fatal neurodegenerative disease that affects nerve cells in the brain and the spinal cord. SURF provides storage and analysis services for the huge amount of DNA data generated by the project.
In het kort
Wie: Jan Veldink en Maarten Kooyman
Functie: arts-hoogleraar ALS Centrum UMC Utrecht en adviseur bij SURF
Organisaties: UMC Utrecht en SURF
Uitdaging: het verwerken en analyseren van enorme hoeveelheden DNA-data (ruim 2 miljoen gigabyte) van duizenden patiënten en controlepersonen vraagt ongekende rekenkracht en veilige opslag.
Oplossing: SURF levert de infrastructuur voor opslag, rekenkracht en data-overdracht.
'I have now died'
Hard-hitting ad campaigns like these have been raising awareness of ALS in recent years. ALS is a progressive condition of the motor nerve cells that mainly affects middle-aged people. “It usually starts with weakness in the arms, legs or speech muscles,” Jan Veldink explains. “That weakness then keeps getting worse and there is no treatment for it. After 3 to 5 years, the respiratory muscles will have weakened to such an extent that people die.”
Veldink is a doctor and professor at the ALS Centre at the University Medical Centre of Utrecht, where around 80% of all Dutch ALS patients receive their treatment. Two of these patients, both entrepreneurs, saw a freezer full of DNA samples at the centre. When they heard that there was no money for further research into the genetic causes of ALS, they offered to help. That was the start of the MinE project.
DNA material from 15,000 patients
“We know that ALS is hereditary,” Veldink says, “but not to such an extent that family research is enough. Therefore, MinE focuses on large-scale DNA study that compares ALS patients with healthy people.”
So how do you obtain that DNA? Large-scale population studies have been conducted with DNA sampling in recent years. However, for a rare disease such as ALS, the entire genome needs to be mapped out in detail, as no one knows which pieces of DNA are important. Material from at least 15,000 patients and 7,500 control subjects is needed. This large control group is crucial because rare genetic variation can differ a lot from country to country – even from region to region within a single country.
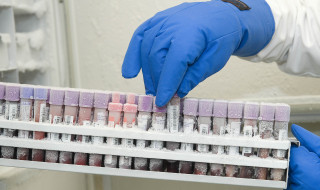
"If you were to analyse the total dataset on a PC, you're in for 600 to 700 years"
The MinE project was set up with the energy you can expect from entrepreneurs. A low-cost contract was concluded with the American company Illumina for the sequencing (reading) of the DNA samples. Crowdfunding campaigns have been set up in each participating country. The project also received funding from the Netherlands ALS Foundation, which attracted many donations thanks to initiatives such as the Ice Bucket Challenge and Amsterdam City Swim.
Veldink: “The Netherlands, Belgium, England, Ireland and the United States are already making significant contributions in terms of fundraising and the supply of samples. Countries such as Italy, Brazil, Australia and France are also well on their way. Finally, Germany and Sweden are doing the sequencing in their own country, but they do share their data with the MinE project.”
Stacks of hard disks as high as a cathedral tower
Raising funds and attracting participants were only the start of the challenges. The sequencing of 1 DNA sample gives you a file of 75 to 100 gigabytes. If you multiply that by 22,500, you get around 2 million gigabytes, which corresponds to a stack of hard drives the height of a cathedral tower. So where do you put all that data? And, more importantly, where do you find the computing power to detect crucial information?
“Many of these types of DNA projects run into barriers in the area of computation,” Veldink says. “Fortunately, we soon came into contact with SURF.” SURF advisor Maarten Kooyman confirms that it is certainly not an easy task. “This huge amount of data is distributed over tens of thousands of separate files. If you were to analyse the total data set on a PC, it would take you between 600 and 700 years. On the Life Science Grid, we can do this in a few weeks using some handy tricks.”
SURF is also working on a direct network connection with Illumina. This means that the data is immediately transferred to the Netherlands as it is read. Veldink: “We prefer to have the genetic data on European soil. So far Illumina has sent this type of data by posting hard disks. That is not convenient and not optimally secure either.”
The network connection is an interesting challenge for SURF. Kooyman: “There will always be bumps in the road, especially with such large quantities. And with such large data packages, the smallest bump becomes a mountain.”
Interesting for many research projects
Veldink is excited about the collaboration with SURF. “It is wonderful that something like SURF exists and that it is practically around the corner from us. It has enabled us to take the lead on this project rather than another institute abroad. It is really special for us that MinE is happening on Dutch soil, in this small country.”
Once the data set is complete – the progress bar now stands at 23% – MinE will have a DNA file that is unique due to its combination of size and quality. The data from the 7,500 healthy control subjects is particularly interesting for many research projects.
“A huge number of sequencing studies are currently being designed,” Veldink says. “They often focus on common diseases such as dementia, autism and schizophrenia. Control groups are important for all kinds of illnesses because you need to look beyond family relationships. The MinE project's data will also be available for that type of research.”
Further reading
- Project MinE
- ALS Centre Netherlands
- Blog Jan Veldink: Largest genetic study in ALS identifies 15 genetic risk factors (9 Dec 2021)
- Blog Jan Veldink: Project MinE update: prestigious prize for innovation in ALS (Feb. 2021)
Author: Aad van de Wijngaard
This article is from SURF Magazine 2015-02
